-Search query
-Search result
Showing all 35 items for (author: wilfling & f)

EMDB-18110: 
The fibrillar and amorphous states of polyQ Q97
Method: electron tomography / : Zhao DY

EMDB-18114: 
phagophore in fibrillar polyQ
Method: electron tomography / : Zhao DY
EMDB-18115: 
phagophore and lysosomes with amorphous polyQ
Method: electron tomography / : Zhao DY

EMDB-19194: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #2
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ
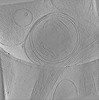

EMDB-19346: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #1
Method: electron tomography / : Hoyer MJ, Capitanio C, Smith IR, Paoli JC, Bieber A, Jiang Y, Paulo JA, Gonzalez-Lozano MA, Baumeister W, Wilfling F, Schulman BA, Harper WJ

EMDB-16417: 
Tomogram of an intracellular Salmonella within a ruptured SCV
Method: electron tomography / : Meijing L

EMDB-16418: 
Tomogram of ruptured SCV with early phagophores
Method: electron tomography / : Meijing L

EMDB-16419: 
Tomogram of stick-shaped contacts between an extended phagophore and the ER around SCV
Method: electron tomography / : Meijing L

EMDB-16420: 
Tomogram of three extended phagophores around SCV
Method: electron tomography / : Meijing L

EMDB-16421: 
Tomogram of omegasomes around SCV
Method: electron tomography / : Meijing L

EMDB-16422: 
Tomogram of membrane contacts between a dilated phagophore rim and the ER around SCV
Method: electron tomography / : Meijing L

EMDB-15526: 
In situ cryo-electron tomogram of a bulk autophagy phagophore in S. cerevisiae
Method: electron tomography / : Bieber A, Capitanio C, Schulman BA, Baumeister W, Wilfling F

EMDB-15545: 
In situ cryo-electron tomogram of a bulk autophagy phagophore in S. cerevisiae #2
Method: electron tomography / : Capitanio C, Bieber A, Erdmann PS, Schulman BA, Baumeister W, Wilfling F

EMDB-15546: 
In situ cryo-electron tomogram of a bulk autophagy autophagosome with END cargo in S. cerevisiae #1
Method: electron tomography / : Bieber A, Capitanio C, Erdmann PS, Schulman BA, Baumeister W, Wilfling F

EMDB-15547: 
In situ cryo-electron tomogram of a bulk autophagy autophagosome fusing with the vacuole in S. cerevisiae #1
Method: electron tomography / : Bieber A, Capitanio C, Erdmann PS, Schulman BA, Baumeister W, Wilfling F

EMDB-15548: 
In situ cryo-electron tomogram of a bulk autophagy phagophore in S. cerevisiae #3
Method: electron tomography / : Bieber A, Capitanio C, Erdmann PS, Schulman BA, Baumeister W, Wilfling F

EMDB-15549: 
In situ cryo-electron tomogram of a bulk autophagy phagophore in S. cerevisiae #4
Method: electron tomography / : Capitanio C, Bieber A, Erdmann PS, Schulman BA, Baumeister W, Wilfling F

EMDB-14323: 
Structure of Chelator-GIDSR4 bound to Mdh2
Method: single particle / : Sherpa D, Chrustowicz J, Schulman B

EMDB-14324: 
Structure of Cage-GIDSR4 bound to PHSVTP-Fbp1
Method: single particle / : Chrustowicz J, Sherpa D, Qiao S, Schulman B

EMDB-14338: 
Structure of endogenous Cage-GIDAnt complex
Method: single particle / : Sherpa D, Chrustowicz J, Qiao S, Schulman B

EMDB-32830: 
GID subcomplex: Gid12 bound Substrate Receptor Scaffolding module
Method: single particle / : Qiao S, Cheng JD, Schulman BA

EMDB-32831: 
Gid12 bound GIDSR4 E3 ubiquitin ligase complex
Method: single particle / : Qiao S, Cheng DJ, Schulman BA
EMDB-32833: 
Gid12 bound Chelator-GIDSR4
Method: single particle / : Qiao S, Cheng DJ, Schulman BA

EMDB-32834: 
Cage assembly GID E3 ubiquitin ligase
Method: single particle / : Qiao S, Cheng DJ, Schulman BA

EMDB-32835: 
Gid12 bound Cage-GIDSR3
Method: single particle / : Qiao S, Cheng DJ, Schulman BA

PDB-7wug: 
GID subcomplex: Gid12 bound Substrate Receptor Scaffolding module
Method: single particle / : Qiao S, Cheng JD, Schulman BA

EMDB-11043: 
In situ cryo-electron tomography reveals layered organization of pre-ribosome maturation in nucleoli. Small Subunit Processome, Class 1
Method: subtomogram averaging / : Erdmann PS, Klumpe S, Hou Z, Beck F, Wilfling F, Plitzko JM, Baumeister W

EMDB-11044: 
In situ cryo-electron tomography reveals layered organization of pre-ribosome maturation in nucleoli. Small Subunit Processome, Class 2
Method: subtomogram averaging / : Erdmann PS, Klumpe S, Hou Z, Beck F, Wilfling F, Plitzko JM, Baumeister W

EMDB-11045: 
In situ cryo-electron tomography reveals layered organization of pre-ribosome maturation in nucleoli. Small Subunit Processome, Class 3
Method: subtomogram averaging / : Erdmann PS, Klumpe S, Hou Z, Beck F, Wilfling F, Plitzko JM, Baumeister W

EMDB-11046: 
In situ cryo-electron tomography reveals layered organization of pre-ribosome maturation in nucleoli. Large Subunit Pre-Ribosome, Class 1
Method: subtomogram averaging / : Erdmann PS, Klumpe S, Hou Z, Beck F, Wilfling F, Plitzko JM, Baumeister W

EMDB-11047: 
In situ cryo-electron tomography reveals layered organization of pre-ribosome maturation in nucleoli. Large Subunit Pre-Ribosome, Class 2
Method: electron tomography / : Erdmann PS, Klumpe S, Hou Z, Beck F, Wilfling F, Plitzko JM, Baumeister W

EMDB-11048: 
In situ cryo-electron tomography reveals layered organization of pre-ribosome maturation in nucleoli. Large Subunit Pre-Ribosome, Class 3
Method: subtomogram averaging / : Erdmann PS, Klumpe S, Hou Z, Beck F, Wilfling F, Plitzko JM, Baumeister W

EMDB-10660: 
Nup116_delta_NPC_25C
Method: subtomogram averaging / : Allegretti M, Zimmerli CE, Rantos V, Wilfling F, Ronchi P, Fung HKH, Lee CW, Hagen W, Turonova B, Karius K, Zhang X, Mulller C, Schwab Y, Mahamid J, Pfander B, Kosinski J, Beck M

EMDB-10661: 
Nup116delta_NPC_37C
Method: subtomogram averaging / : Allegretti M, Zimmerli CE, Rantos V, Wilfling F, Ronchi P, Fung HKH, Lee CW, Hagen W, Turonova B, Karius K, Zhang X, Muller C, Schwab Y, Mahamid J, Pfander B, Kosinski J, Beck M

EMDB-10198: 
In-cell S cerevisiae nuclear pore complex
Method: subtomogram averaging / : Beck M, Allegretti M
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
